The International Space station's Surface Microbial Profile Resembles Skin Of Its Crew Members

A study conducted by a team of national laboratory and NASA researchers has found that the environment of the International Space Station is affected by the microbial composition of the astronauts themselves.
The five-year research effort represents the first study to compare the space station’s environmental microbial profile (or microbiome) to an astronaut’s microbiome using metagenomic DNA sequencing techniques.
Their paper, published yesterday in the scientific journal Public Library of Science (PLOS) One, is the work of scientists from Lawrence Livermore National Laboratory (LLNL) and three NASA centers – the Jet Propulsion Laboratory (JPL), Ames Research Center in Silicon Valley and Johnson Space Center.
“It’s very important to have continual monitoring of the microbiome on the space station because it will help us identify the potential for any microbes that could harm the astronauts’ health,” said LLNL biologist Crystal Jaing, the paper’s lead author and the principal investigator for the Microbial Tracking (MT)-2 study.
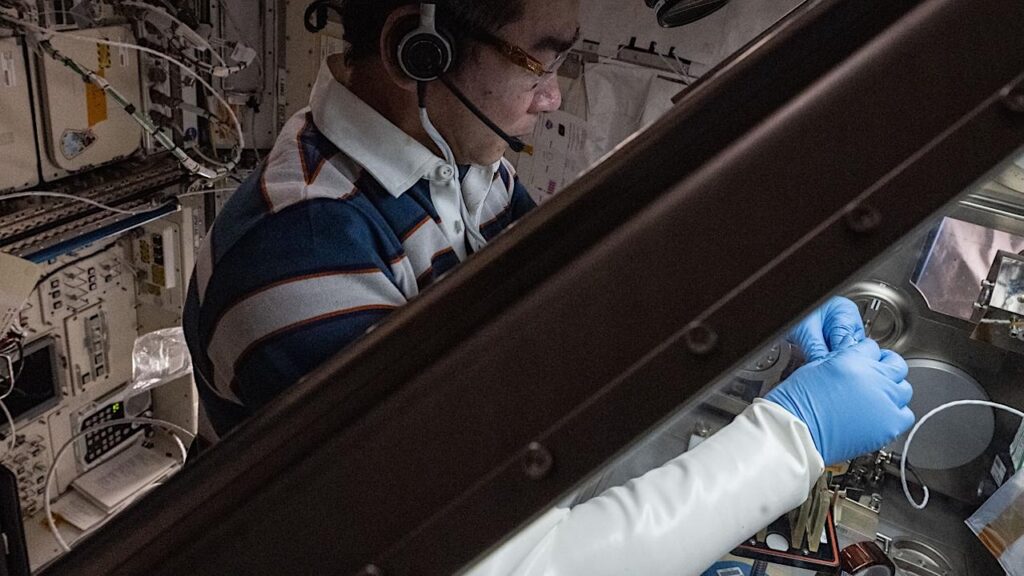
The scientists characterized one astronaut’s microbial profile, taking 88 body swabs of the astronaut’s mouth, nose, ear, skin and saliva – and found that the microbiome of space station surfaces resembled the crew member’s skin.
The astronaut samples were collected at eight different times – before the flight (22), during the flight (33) and post flight (33).
To understand if the crew microbiome interacted with the space station’s environment, environmental surface samples from eight different habitable locations in the space station were collected from two flights. Environmental samples from one flight were collected by the astronaut whose microbial profile was compiled and samples from the next flight were collected after the astronaut departed.
The microbial composition of samples from both the astronaut and the space station’s environment was measured using metagenomic sequencing and processed using the Livermore Metagenomics Analysis Toolkit, a bioinformatics software that rapidly identifies microbes from huge amounts of DNA sequence data.
Although this study used samples returned from space, NASA has the ability to identify microbes in real time aboard the space station and is planning real time microbial monitoring on future spacecrafts as well.

Top 12 most abundant species from crewmember samples pre-, during- and post-flight. The percent of mapped reads from saliva, mouth, nostril, ear and skin samples for each species in each sample (ranked by the average abundance in each panel summed across locations). Each sample’s time point label (day code) and day number are shown.
This study builds upon the Microbial Tracking-1 project, in which the microbial diversity from space station surfaces and air samples were characterized by conventional microbiology but not correlated with the astronauts’ microbiome. The MT-2 study expanded upon this study by collecting parallel astronaut samples, along with samples from the exact same surfaces studied by MT-1.
“We found in this study that microbial populations of space station surfaces resemble those of the crew’s skin. This might aid in designing preventive measures to minimize risk against potentially problematic microorganisms onboard spacecraft destined for long-term space travel,” said Kasthuri Venkateswaran, a microbiologist with NASA’s JPL and one of the paper’s co-authors.
The researchers particularly want to understand what changes happen in the space station’s microbial communities during space flight.
“Our main purpose is to understand the microbial communities in the space station, so we can use that information to design diagnostics and countermeasures to ensure crew health, particularly for long space flights, such as to the moon and Mars,” Jaing said.
Jaing’s colleague and PLOS ONE paper co-author, NASA Ames microbiologist David Smith, agreed, adding that human beings carry microorganisms everywhere they travel, including space.
“The long-term measurements from this study will provide insights about which spacecraft-associated microbes to expect on a long-duration human mission,” Smith said.
More study of the impact of microbes on humans in closed environments is needed, the researchers said, as “the beneficial, benign or detrimental pathways associated with microbe-host interactions remain poorly understood both on Earth and in space.”
Astronaut health is protected through routine monitoring of microbes on the space station, along with good nutrition and appropriate exercise, as well as good personal and space station sanitation processes.
“People with uncompromised respiratory tract and healthy immune systems can usually fight off potentially harmful microbes residing within built habitats. However, human spaceflight introduces new risks that must be considered because astronauts can become more susceptible to infections during long spaceflight missions,” the scientists wrote.
In their paper, the researchers observe that the space station, an “island” in space, offers “an unparalleled location” for studying the interactive dynamics between humans and microorganisms inside closed habitats.
In addition to Jaing, Venkateswaran and Smith, other team members are biologists Nick Be and James Thissen and computer scientist Aram Avila Herrera, all of LLNL; microbiologist Camilla Urbaniak of NASA-JPL; space biologist Fathi Karouia of Blue Marble Space at NASA Ames; and virologist Satish Mehta of Wyle Laboratories Inc. at NASA’s Johnson Space Center.
The paper is the first of several studies that will be published with new information by the research team that could help protect not only astronauts on space exploration missions but also people on Earth.
NASA’s Space Life and Physical Sciences Research and Applications Division funded the MT-1 and MT-2 studies.
Crewmember microbiome may influence microbial composition of ISS habitable surfaces PLOS One
Astrobiology