Genome Modeling And Design Across All Domains Of Life With Evo 2
All of life encodes information with DNA. While tools for sequencing, synthesis, and editing of genomic code have transformed biological research, intelligently composing new biological systems would also require a deep understanding of the immense complexity encoded by genomes.
See With Evo 2, AI Can Model And Design The Genetic Code For All Domains Of Life, Nature 7 March 2026
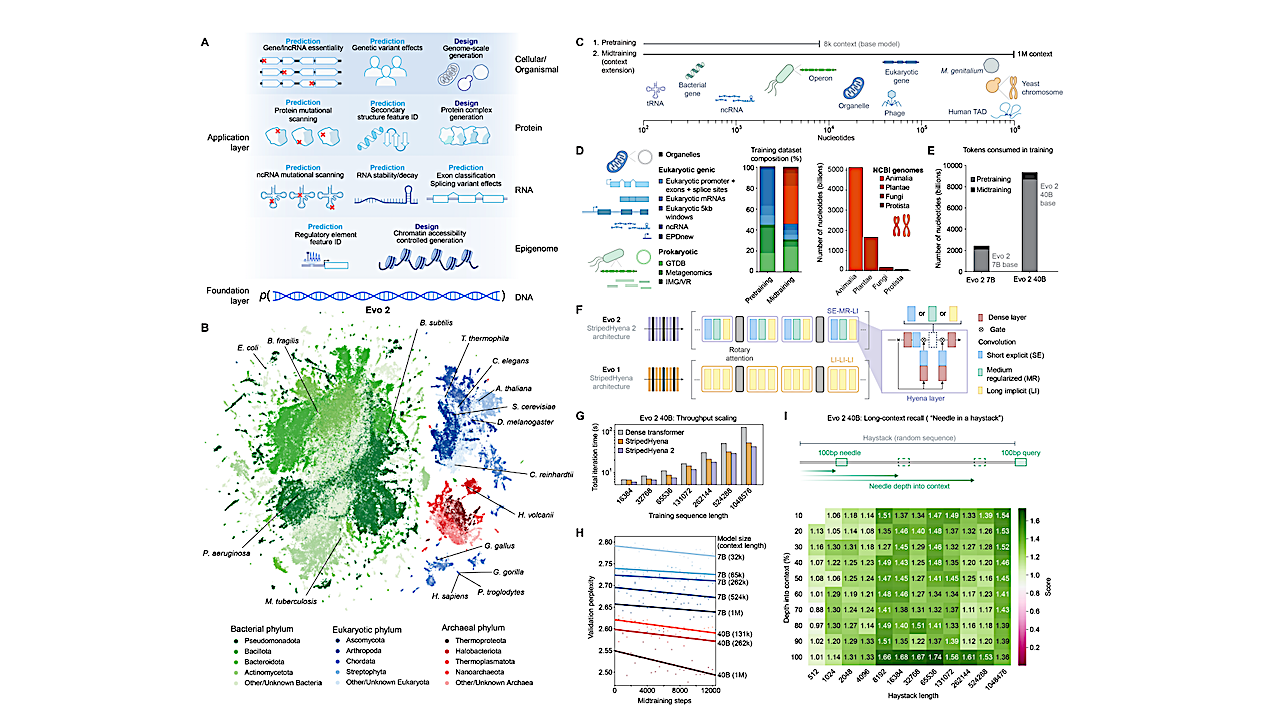
We introduce Evo 2, a biological foundation model trained on 9.3 trillion DNA base pairs from a highly curated genomic atlas spanning all domains of life. We train Evo 2 with 7B and 40B parameters to have an unprecedented 1 million token context window with single-nucleotide resolution. Evo 2 learns from DNA sequence alone to accurately predict the functional impacts of genetic variation—from noncoding pathogenic mutations to clinically significant BRCA1 variants—without task-specific finetuning.
Applying mechanistic interpretability analyses, we reveal that Evo 2 autonomously learns a breadth of biological features, including exon–intron boundaries, transcription factor binding sites, protein structural elements, and prophage genomic regions. Beyond its predictive capabilities, Evo 2 generates mitochondrial, prokaryotic, and eukaryotic sequences at genome scale with greater naturalness and coherence than previous methods.
Guiding Evo 2 via inference-time search enables controllable generation of epigenomic structure, for which we demonstrate the first inference-time scaling results in biology.
We make Evo 2 fully open, including model parameters, training code, inference code, and the OpenGenome2 dataset, to accelerate the exploration and design of biological complexity.
Genome modeling and design across all domains of life with Evo 2, biorxiv.org
See With Evo 2, AI Can Model And Design The Genetic Code For All Domains Of Life, Nature 7 March 2026
Astrobiology, genomics, omics, bioinformatics, evolution,