Modularity, Ecology, And Theoretical Evolution Of The Ribozyme Body Plan
Ribozymes are relics of molecular life forms from the primitive earth that are embedded within modern genomes across all kingdoms of life. Despite significant knowledge from decades of bioinformatic and biochemical research, a gap remains in our understanding of the world in which ribozymes existed, their interactions, ecology, and possibly also evolution.
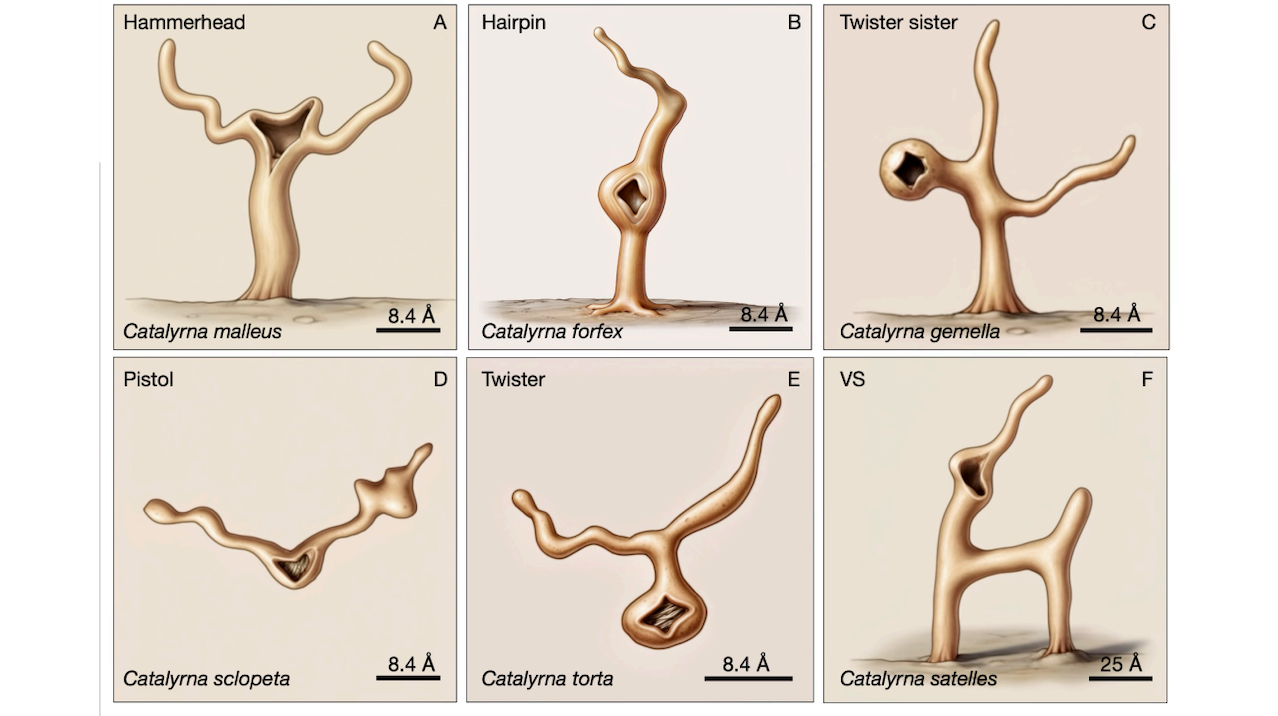
The present study proposes a new theoretical basis for understanding these aspects of ribozyme biology by adopting a zoological frame of thought. Seven families of small self-cleaving ribozymes are each mapped to a primitive marine animal analog based on topological architecture, and classified into body plan grades paralleling cnidarian, ctenophore, and bilaterian organization.
A formal notation describing ribozyme regions as bodies, cavities, and limbs enables systematic comparison with animal body plans and highlights reusability of parts across ribozyme groups, in turn enabling the construction of a connectivity network and a putative body plan-based evolutionary ordering.
This ordering of body plans identifies systematic gaps corresponding to undiscovered ribozyme forms, one of which, a planktonic form of hammerhead, was bioinformatically found in 16.2% of all hammerhead sequences. Computational cross-cleavage analysis across all 49 pairwise interactions (including conspecific) suggests that the hammerhead was a generalist apex predator in the RNA world, while the hatchet was a vulnerable, filter-feeding or scavenger prey species. Conspecific analysis suggests that cannibalism was also a prevalent feeding strategy.
Evolutionary avoidance signatures suggest ancient predator-prey coevolution. This theory emphasizes behavior, modularity, and ecological interactions as primary drivers of early ribozyme evolution, offering a new pathway for inferring ancient RNA forms independent of sequence-first assumptions.
Modularity, ecology, and theoretical evolution of the ribozyme body plan, biorxiv.org
Astrobiology, genomics, evolution,