Oxygen Reductase Origin Followed The Great Oxidation Event And Terminated The Lomagundi Excursion
The history of Earth’s atmospheric oxygen is a cornerstone of evolutionary biology. While unequivocal evidence for an increase in atmospheric O2 marks the Great Oxidation Event (GOE) roughly 2.4 billion years ago, evidence underlying proposals for pre-GOE O2 accumulation is debated.
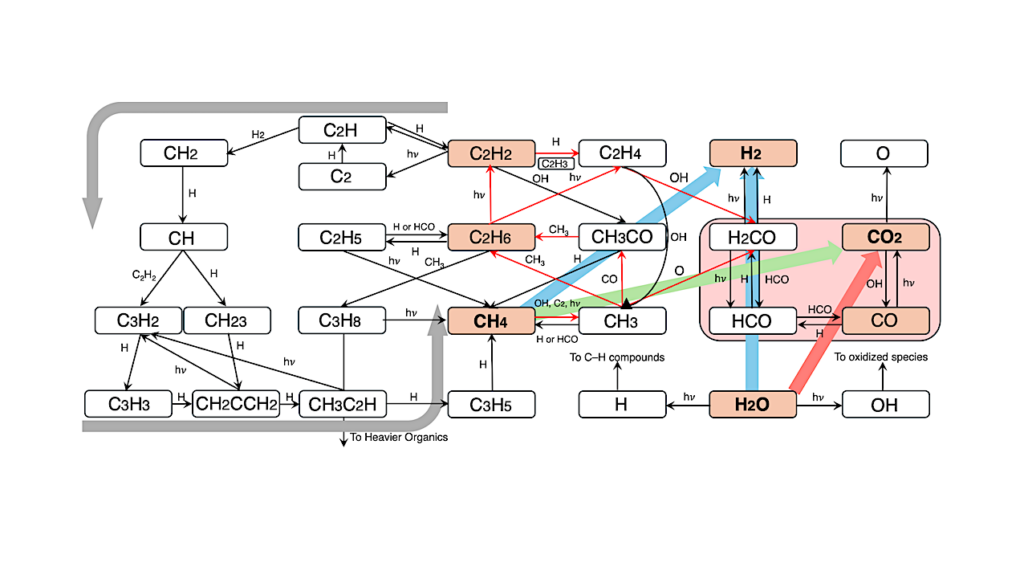
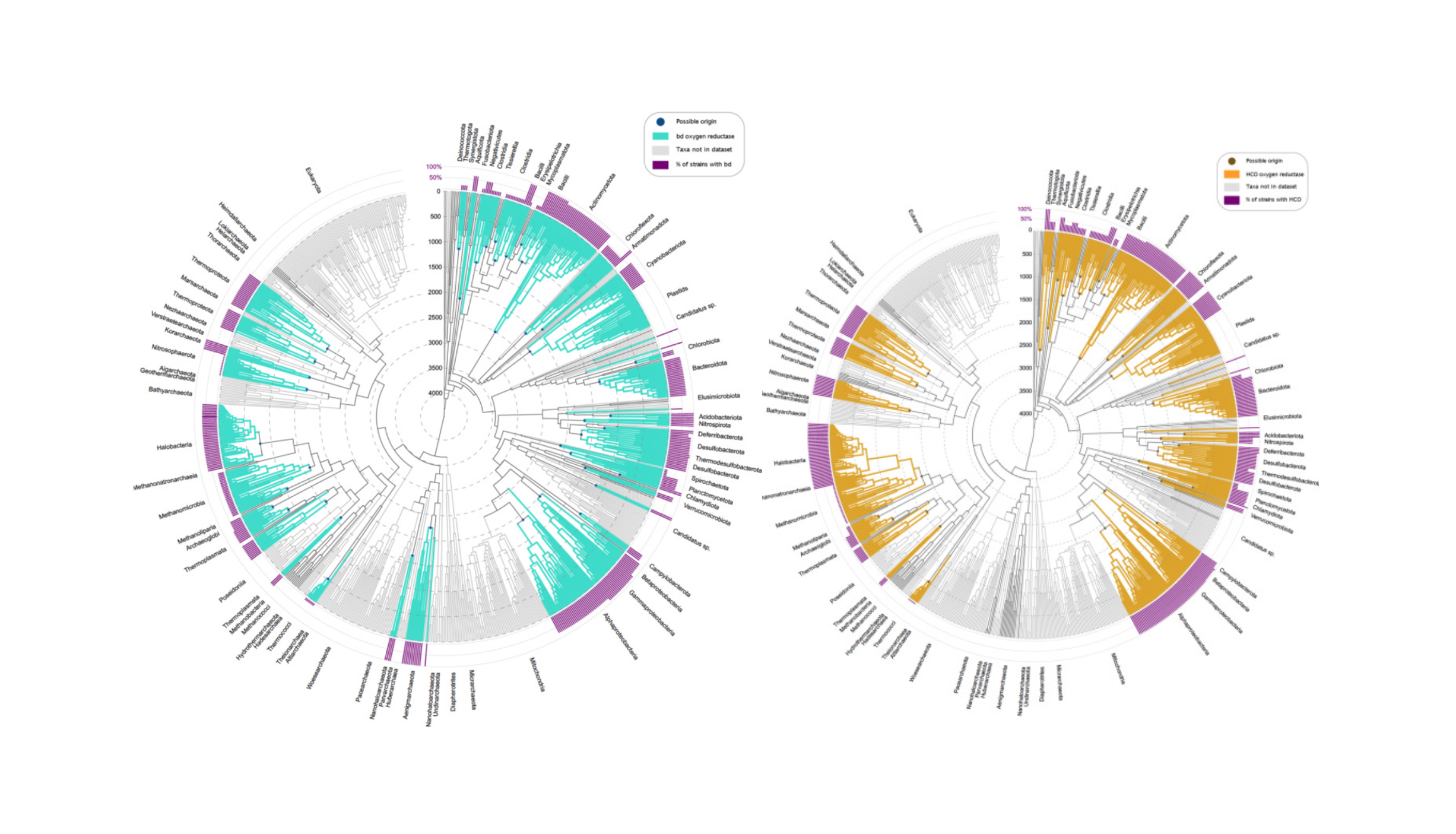
Here we have investigated the distribution of genes for oxygen reductases, the enzymes that consume O2 in respiratory chains, across independently generated molecular timescales of prokaryotic evolution.
The data indicate that cytochrome bd-oxidases, heme-copper oxidases and alternative oxidases arose in the wake of the GOE ca. 2.4 billion years ago, after which the genes were subjected to abundant lateral gene transfer, a reflection of their utility in redox balance and membrane bioenergetics.
The data lead us to propose a straightforward four-stage model for O2 accumulation surrounding the GOE: (i) Negligible O2 existed prior to the GOE. (ii) Cyanobacterial O2 production started at the GOE, yet was capped at 2% [v/v] atmospheric O2, the threshold at which cyanobacterial nitrogenase is inhibited by O2. (iii) Production of 0.02 atm of O22 (2 % [v/v]) at the GOE buried roughly the entire atmospheric CO2 inventory, causing sudden enrichment of 13C in dissolved inorganic carbon (the Lomagundi 13C anomaly), through RuBisCO isotope discrimination, without atmospheric O22 exceeding 2% [v/v]. (iv) High atmospheric 12C at the end of the Lomagundi excursion marks the origin of oxygen reductases, their rapid spread via function in respiratory CO2 liberation, and the onset of equilibrium between photosynthetic O2 production and respiratory O2 consumption at 2 % atmospheric O2.
- Oxygen reductase origin followed the great oxidation event and terminated the Lomagundi excursionm Biochimica et Biophysica Acta (BBA) – Bioenergetics via PubMed
- Oxygen reductase origin followed the great oxidation event and terminated the Lomagundi excursion, Biochimica et Biophysica Acta (BBA) (open access)
Astrobiology,