Model Suggests How Ancient RNA May Have Gained Self-Cutting Ability Essential for Life

Scientists have long pondered the beginnings of life on Earth. One theory is that RNA, which is ubiquitous across all domains of life, played a central role in early life. Similar to DNA, RNA possesses the ability to store genetic information. However, to initiate life’s processes, early RNA must have also possessed the capability to self-replicate and catalyze biochemical reactions independently, without the assistance of specialized enzymes.
Previously, it was unclear how a molecule this complex could arise without any precursors. However, in a new study published in eLife, Alexei Tkachenko, a physicist at Brookhaven National Laboratory, and Sergei Maslov (CAIM co-leader/CABBI), a professor of bioengineering and physics at the University of Illinois Urbana-Champaign, describe their model that demonstrates how such a molecule could gain functionality.
“RNA is very preserved across all organisms. So, it’s like a smoking gun that RNA has a central point in life,” explained Tkachenko. “The problem is that modern RNA is very finely tuned and complex, so we wanted to see how life might emerge from something much simpler.”
While modern RNA relies on enzymes for replication and catalytic activity, the origins of these enzymes themselves remain a puzzle. However, the discovery of ribozymes, RNA molecules exhibiting enzymatic properties, suggests a plausible pathway for the emergence of early functional polymers.
The challenge lies in understanding how these ancient RNA molecules could have possessed the ability to “cut” other molecules, a crucial step in the replication process of DNA and RNA.
“Experiments have shown that a cleavage ribozyme, which relies on only a handful of conserved bases to do its job, can emerge spontaneously with no prior information, if the experimenter artificially selects for things that cut other things,” said Tkachenko. “The problem is that it’s not clear how evolution would select for something that cuts things, essentially selecting for a destructive enzyme.”
To tackle this question, the researchers devised a model simulating basic RNA molecules devoid of enzymatic activity. Within this model, random bond breakage was allowed to occur, mimicking real-world chemical processes. The researchers observed that breakage led to more copies of the polymer that was broken, meaning that molecules capable of self-cleavage would have been favored by evolution due to their ability to replicate.
Maslov illustrated this concept with an analogy, likening the process to cutting an earthworm in half, where both halves regenerate into whole organisms.
“In principle, if you wanted to make many earthworms from one earthworm, you would just start cutting them one by one,” explained Maslov. “The same idea is why cutting would be selected for in RNA, because when it’s cut it regrows itself from individual building blocks. And that was the connection, to explain why the first ribozyme was selected to cut things — because cutting is how RNA exponentially grows.”
But how does catalytic activity arise from such simple beginnings? In a second model, the researchers demonstrated how RNA molecules could evolve into complex ecosystems with functional properties, where different polymers in these ecosystems cleave and replicate each other.
Their model simulated a pool of polymer chains competing for nucleotide “building blocks”, and cutting other polymers they encountered. Polymer chains pair in specific ways (such as the A-T nucleotide pairing in modern DNA), and as such, the chains in the simulation formed template and complimentary strands, essentially working together.
“Pairing rules are the basis for how information is preserved and propagated in the future,” said Maslov. “And it’s also important for function, because it gives way to hairpins in the strands that lead to a three-dimensional shape, and these are the ones which are capable of enzymatic activity.”
Polymer replication in the model occurred based on temperature being cycled between hot and cold phases (typical in day-night cycles), suggesting that ancient polymers may have relied on such cycles to grow. Nonorganic surfaces such as rocks may have also facilitated this process.
These findings offer compelling insights into the natural emergence and selection of ribozymes with enzymatic activity, shedding light on a crucial aspect of early life evolution. The researchers advocate for experimental validation of their models to confirm their predictions.
Additionally, they acknowledge the discrepancy between bidirectional growth in their model and the unidirectional growth observed in DNA and RNA replication in real life. Alexei says they plan to continue adjusting the model to see if they can find variations which would result in unidirectional growth.
“It is not a coincidence that Carl Woese, who our genomic institute is named after, used pieces of ribosomal RNA to make his trees of life,” said Maslov. “RNA inside ribosomes is universal to every single organism from bacteria to archaea to eukaryotes like you and me. This paper definitely doesn’t solve the problem of origin of life, but it fills a tiny gap in our understanding of how early RNA may have functioned to bring about life.”
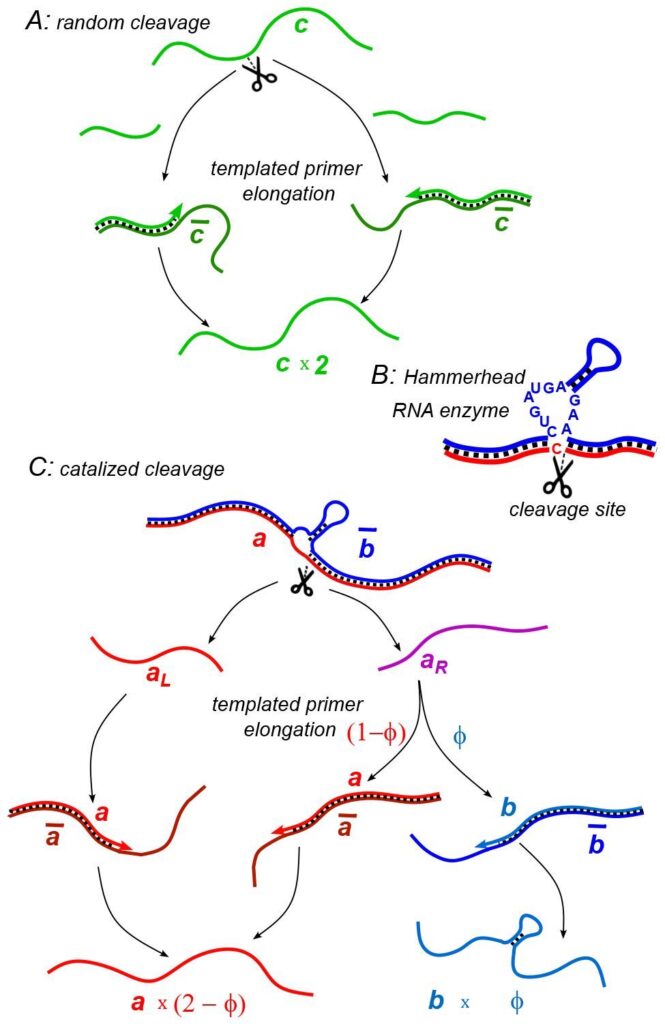
(A) Random cleavage model. A random break in a chain of type c generates two primer fragments, which are elongated to give rise to two chains of type c. Elongation requires a complementary template of type . (B) An example of catalyzed cleavage given by hammerhead ribozyme [33]. Note that the right cleavage fragment is perfectly complementary to the blue sequence, while the left one contains an extra non-complementary base C. (C) Catalyzed cleavage model. A cleavage of the red chain a catalyzed by the blue chain gives rise to two primers aL (red) and aR (purple). Because of an extra non-complementary base (see Panel B) the aL primer can only elongate to a, while the aR primer – to either a or b depending on its first hybridization partner. Similar processes involving complementary chains ā and (not shown) result in the replication of templates.
This study was partially funded by U.S. Department of Energy Office of Science and can be found at https://doi.org/10.7554/eLife.91397.2
Emergence of catalytic function in prebiotic information-coding polymers, elife (open access)
By Shelby Lawson
This news release was originally issued by the University of Illinois Urbana-Champaign. Alexei Tkachenko is a staff scientist in the Theory and Computation group at the Center for Functional Nanomaterials (CFN), a U.S. Department of Energy (DOE) Office of Science user facility at DOE’s Brookhaven National Laboratory. For more information of Brookhaven’s role in this work, contact Stephanie Kossman ([email protected], 631-344-8671).
astrobiology