How Dormant Bacteria Come Back To Life
Solving a riddle that has confounded biologists since bacterial spores — inert, sleeping bacteria — were first described more than 150 years ago, researchers at Harvard Medical School have discovered a new kind of cellular sensor that allows spores to detect the presence of nutrients in their environment and quickly spring back to life.
It turns out that these sensors double as channels through the membrane and remain closed during dormancy but rapidly open when they detect nutrients. Once open, the channels allow electrically charged ions to flow out through the cell membrane, setting in motion the shedding of protective spore layers and the switching on of metabolic processes after years — or even centuries — of dormancy.
The team’s findings, published April 28 in Science, could help inform the design of ways to prevent dangerous bacterial spores from lying dormant for months, even years, before waking up again and causing outbreaks.
“This discovery solves a puzzle that’s more than a century old,” said study senior author David Rudner, professor of microbiology in the Blavatnik Institute at HMS. “How do bacteria sense changes in their environment and take action to break out of dormancy when their systems are almost completely shut down inside a protective casing?”
How sleeping bacteria come back to life
To survive adverse environmental conditions, some bacteria go into dormancy and become spores, with biological processes put on hold and layers of protective armor around the cell.
These biologically inert mini fortresses allow bacteria to wait out periods of famine and shield themselves from the ravages of extreme heat, dry spells, UV radiation, harsh chemicals, and antibiotics.
For more than a century, scientists have known that when the spores detect nutrients in their environment, they rapidly shed their protective layers and reignite their metabolic engines. Although the sensor that enables them to detect nutrients was discovered almost 50 years ago, the means of delivering the wake-up signal, and how that signal triggers bacterial revival remained a mystery.
In most cases, signaling relies on metabolic activity and often involves genes encoding proteins to make specific signaling molecules. However, these processes are all shut off inside a dormant bacterium, raising the question of how the signal induces the sleeping bacteria to wake up.
In this study, Rudner and team discovered that the nutrient sensor itself assembles into a conduit that opens the cell back up for business. In response to nutrients, the conduit, a membrane channel, opens, allowing ions to escape from the spore interior. This initiates a cascade of reactions that allow the dormant cell to shed its protective armor and resume growth.
The scientists used multiple avenues to follow the twists and turns of the mystery. They deployed artificial intelligence tools to predict the structure of the intricately folded sensor complex, a structure made of five copies of the same sensor protein. They applied machine learning to identify interactions between subunits that make up the channel. They also used gene-editing techniques to induce bacteria to produce mutant sensors as a way to test how the computer-based predictions played out in living cells.
“The thing that I love about science is when you make a discovery and suddenly all these disparate observations that don’t make sense suddenly fall into place,” Rudner said. “It’s like you’re working on a puzzle, and you find where one piece goes and suddenly you can fit six more pieces very quickly.”
Rudner described the process of discovery in this case as a series of confounding observations that slowly took shape, thanks to a team of researchers with diverse perspectives working together synergistically.
Along the way, they kept making surprising observations that confused them, hints that suggested answers that didn’t seem like they could possibly be true.
Stitching the clues together
One early clue emerged when Yongqiang Gao, an HMS research fellow in the Rudner lab, was conducting a series of experiments with the microbe Bacillus subtilis, commonly found in soil and a cousin to the bacterium that causes anthrax. Gao introduced genes from other bacteria that form spores into B. subtilis to explore the idea that the mismatched proteins produced would interfere with germination. Much to his surprise, Gao found that in some cases the bacterial spores reawakened flawlessly with a set of proteins from a distantly related bacterium.
Lior Artzi, a postdoctoral fellow in the lab at the time of this research, came up with an explanation for Gao’s finding. What if the sensor was a kind of receptor that acts like a closed gate until it detects a signal, in this case a nutrient like a sugar or an amino acid? Once the sensor binds to the nutrient, the gate pops open, allowing ions to flow out of the spore.
In other words, the proteins from distantly related bacteria would not need to interact with mismatched B. subtilis spore proteins, but instead simply respond to changes in the electric state of the spore as ions begin to flow.
Rudner was initially skeptical of this hypothesis because the receptor didn’t fit the profile. It had almost none of the characteristics of an ion channel. But Artzi argued the sensor might be made up of multiple copies of the subunit working together in a more complex structure.
AI has entered the chat
Another postdoc, Jeremy Amon, an early adopter of AlphaFold, an AI tool that can predict the structure of proteins and protein complexes, was also studying spore germination and was primed to investigate the nutrient sensor.
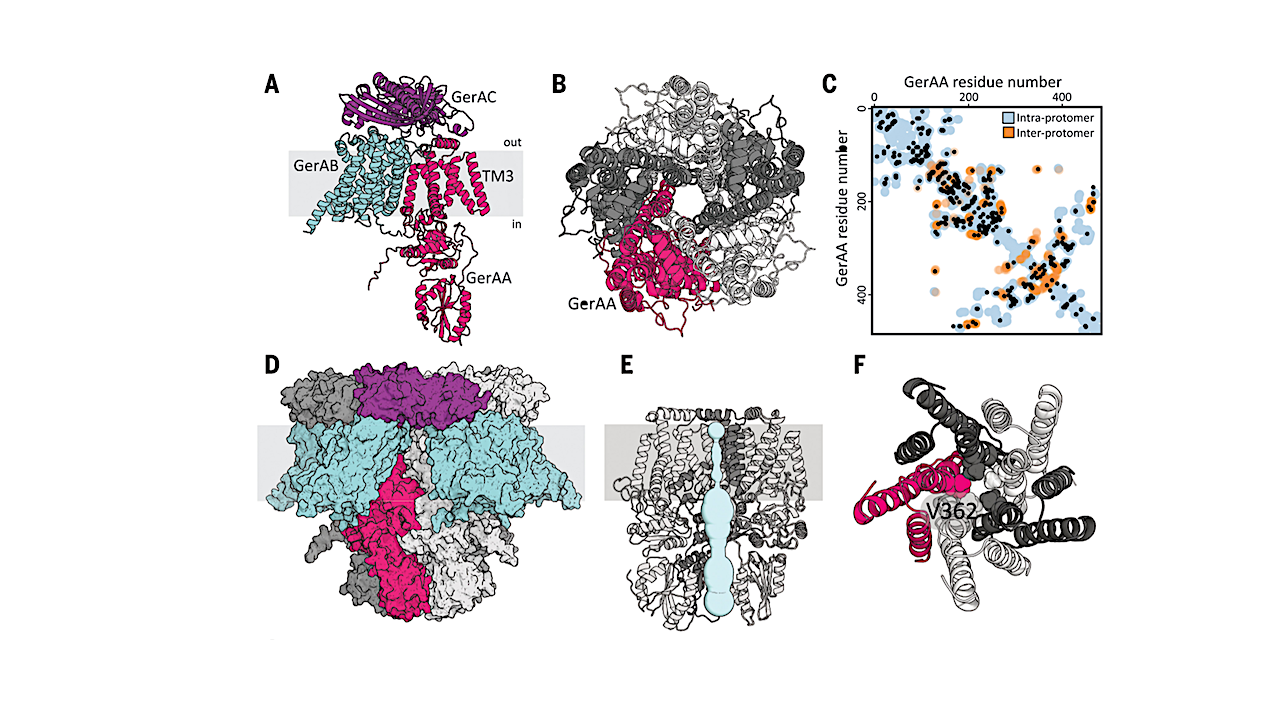
The tool predicted that a particular receptor subunit assembles into a five-unit ring known as a pentamer. The predicted structure included a channel down the middle that could allow ions to pass through the spore’s membrane. The AI tool’s prediction was just what Artzi had suspected.
Gao, Artzi, and Amon then teamed up to test the AI-generated model. They worked closely with a third postdoc, Fernando Ramírez-Guadiana and the groups of Andrew Kruse, HMS professor of biological chemistry and molecular pharmacology, and computational biologist Deborah Marks, HMS associate professor of systems biology.
They engineered spores with altered receptor subunits predicted to widen the membrane channel and found the spores awoke in the absence of nutrient signals. On the flip side, they generated mutant subunits that they predicted would narrow the channel aperture. These spores failed to open the gate to release ions and awake from stasis in the presence of ample nutrients to coax them out of dormancy.
In other words, a slight deviation from the predicted configuration of the folded complex could leave the gate stuck open or shut, rendering it useless as a tool for waking up the dormant bacteria.
Implications for human health and food safety
Understanding how dormant bacteria spring back into life is not just an intellectually tantalizing puzzle, Rudner said, but one with important implications for human health. A number of bacteria that are capable of going into deep dormancy for stretches of time are dangerous, even deadly pathogens: The powdery white form of weaponized anthrax is a made up of bacterial spores.
Another dangerous spore-forming pathogen is Clostridioides difficile, which causes life-threatening diarrhea and colitis. Illness from C. difficile typically occurs after use of antibiotics that kill many intestinal bacteria but are useless against dormant spores. After treatment, C. difficile awakens from dormancy and can bloom, often with catastrophic consequences.
Eradicating spores is also a central challenge in food-processing plants because the dormant bacteria can resist sterilization due to their protective armor and dehydrated state. If sterilization is unsuccessful, germination and growth can cause serious foodborne illness and massive financial losses.
Understanding how spores sense nutrients and rapidly exit dormancy can enable researchers to develop ways to trigger germination early, making it possible to sterilize the bacteria, or block germination, keeping the bacteria trapped inside their protective shells, unable to grow, reproduce, and spoil food or cause disease.
Authorship, funding, disclosures
Additional authors include Kelly Brock and Joshua Cofsky, of HMS.
Support for this work comes from the National Institutes of Health grants GM086466, GM127399, GM122512, AI171308 (DZR), AI164647 (DZR, ACK, DSM) and funds from the Harvard Medical School Dean’s Initiative. Amon was funded by National Institutes of Health grant F32GM130003. Artzi was a Simons Foundation fellow of the Life Sciences Research Foundation.
Bacterial spore germination receptors are nutrient-gated ion channels, Science (open access)
Astrobiology