Photosynthetic Secrets Come To Light
Secrets of photosynthesis have been discovered at atomic level, shedding important new light on this plant super-power that greened the earth more than a billion years ago.
John Innes Centre researchers used an advanced microscopy method called cryo-EM to explore how the photosynthetic proteins are made.
The study, published in Cell, presents a model and resources to stimulate further fundamental discoveries in this field and assist longer term goals of developing more resilient crops.
Dr Michael Webster, group leader and co-author of the paper said: “Transcription of chloroplast genes is a fundamental step in making the photosynthetic proteins that provide plants with the energy they need to grow. We hope that by understanding this process better – at the detailed molecular level – we will equip researchers looking to develop plants with more robust photosynthetic activity.”
“The most important outcome of this work is the creation of a useful resource. Researchers can download our atomic model of the chloroplast polymerase and use it to produce their own hypotheses of how it might function and experimental strategies that would test them.”
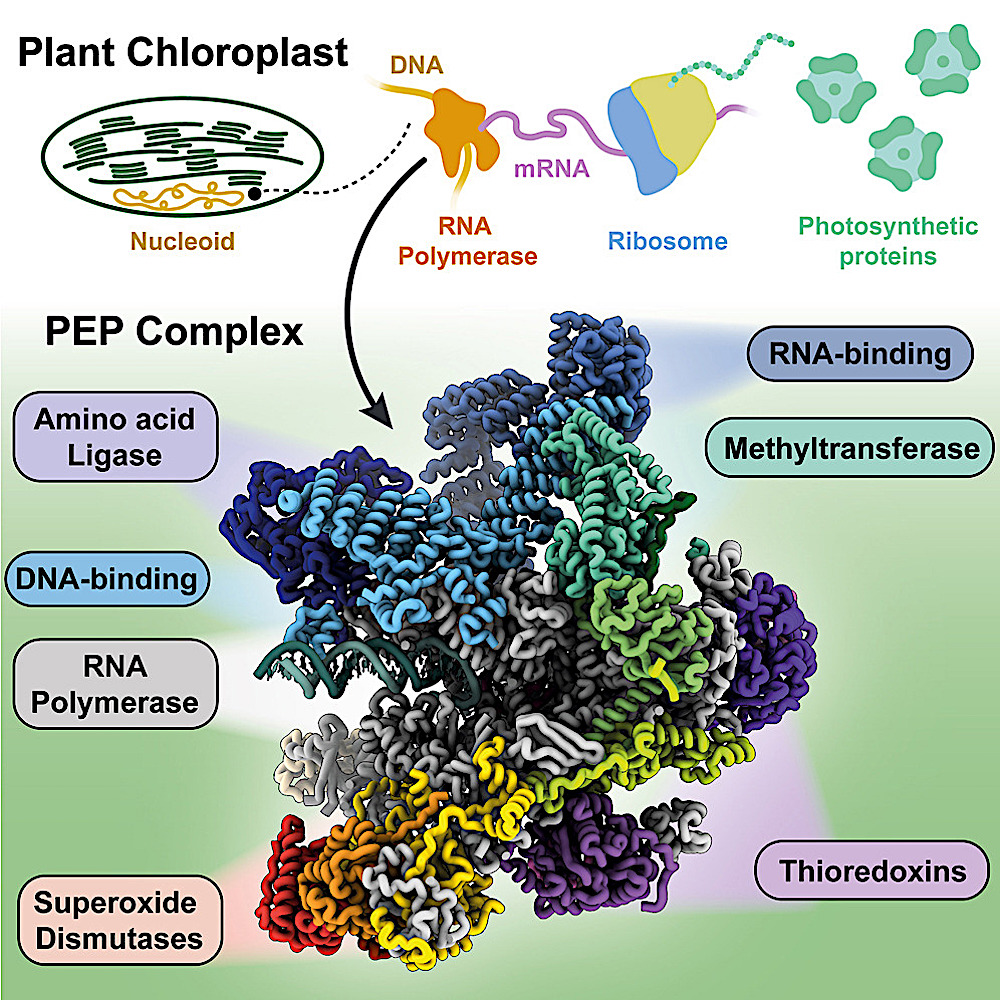
Graphical abstract
Photosynthesis takes place inside chloroplasts, small compartments within plant cells that contain their own genome, reflecting their past as free-living photosynthetic bacteria before they were engulfed and co-opted by plants.
The Webster group at the John Innes Centre investigates how plants make photosynthetic proteins, the molecular machines that make this elegant chemical reaction happen, converting atmospheric carbon dioxide and water into simple sugars and producing oxygen as a byproduct.
The first stage in protein production is transcription, where a gene is read to produce a ‘messenger RNA’. This transcription process is done by an enzyme called RNA polymerase.
It was discovered 50 years ago that chloroplasts contain their own unique RNA polymerase. Since then, scientists have been surprised how complex this enzyme is. It has more subunits than its ancestor, the bacterial RNA polymerase, and is even bigger than human RNA polymerases.
The Webster group wanted to understand why chloroplasts have such a sophisticated RNA polymerase. To do this they needed to visualise the structural architecture of the chloroplast RNA polymerase.
The research team used a method called cryogenic electron microscopy (cryo-EM) to image samples of chloroplast RNA polymerase purified from white mustard plants.
By processing these images, they were able to build a model that contains the positions of more than 50,000 atoms in the molecular complex.
The RNA polymerase complex comprises 21 subunits encoded in the two genomes, nuclear and chloroplast. Close analysis of this structure as it performs transcription allowed the researchers to start explaining these components’ functions.
The model allowed them to identify a protein that interacts with the DNA as it is being transcribed and guides it to the enzyme’s active site.
In this video we summarise our research into the molecular structure of the chloroplast transcription machinery.
Another component can interact with the mRNA that is being produced that likely protects it from proteins that would degrade it before it is translated into protein.
Dr Webster said: “We know that each component of the chloroplast RNA polymerase has a vital role because plants that lack any one of them cannot make photosynthetic proteins and consequently cannot turn green. We are studying the atomic models carefully to pinpoint what the role is for each of the 21 components of the assembly.”
Joint first author Dr Ángel Vergara-Cruces said: “Now that we have a structural model the next step is to confirm the role of the chloroplast transcription proteins. By revealing mechanisms of chloroplast transcription, our study offers insight into its role in plant growth and adaptation and response to environmental conditions.”
Joint first author Dr Ishika Pramanick said: “There were many surprising moments in this remarkable work journey, starting with the very challenging protein purification to taking stunning cryo-EM images of this huge complex protein to finally seeing our work in a printed version.”
Dr Webster concluded: “Heat, drought, and salinity limit a plants’ ability to perform photosynthesis. Plants that can produce photosynthetic proteins reliably in the face of environmental stress may control chloroplast transcription differently. We look forward to seeing our work used in the important effort to develop more robust crops.”
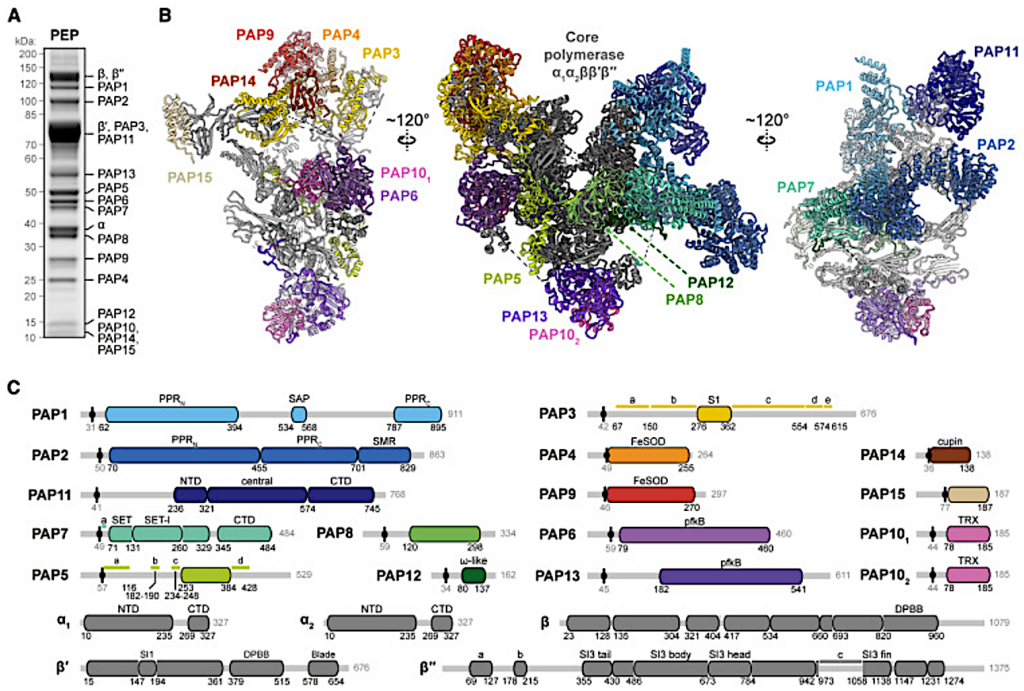
(A) SDS-PAGE of PEP purified from S. alba chloroplasts with identified subunits indicated.
(B) Structural model of PEP (center) and clipped views of the β and β′ lobes (left and right, respectively).
(C) Domain organization of S. alba PEP subunits determined from the structural model. Positions of chloroplast target peptide cleavage sites are indicated with black circle and line.
Alternative protein name of PAP13 is FLN2; alternative of PAP14 is pTAC18; alternative of PAP15 is PRIN2. Additional sequence annotations: PAP3: a, β′′-SI3 fin binding region; b, S1 case; c, arch bridge; d, β binding region; e, β′′-SI3 body binding region; PAP5: a, PIR1; b, PAP8 binding motif; c, rim helix binding motif; d, β binding region; PAP7: a, α-CTD binding motif; β′′: a, rim helices; b, bridge helix; c, S1 case; PAP4 and PAP9: FeSOD, iron superoxide dismutase; PAP6 and PAP13: pfkB, pfkB family carbohydrate kinase domain; PAP10: TRX, thioredoxin. See also Figures S1–S3 and Tables S1–S5.
Structure of the plant plastid-encoded RNA polymerase, Cell (open access)
Astrobiology, Genomics,