New Dimensions Of Extremophile Genomes Discovered
It has long been thought that an organism’s DNA provides clues only about its ancestry and how the various forms of life on Earth are related – the more similar their DNA, the more closely the species are related.
However, using machine learning, researchers at the University of Waterloo have discovered that the DNA of some species called “extremophiles” living in comparably harsh environments is strikingly similar despite these organisms being only very distantly related.
“At first, we could not believe our eyes, as it was so unexpected,” said Dr. Lila Kari, a professor at the Cheriton School of Computer Science at the University of Waterloo. “This is in a way akin to finding out that your human DNA is more similar to the plant DNA of flowers in your garden, than to the human DNA of your cousin who lives on another continent.”
Extremophiles are organisms that not only inhabit but thrive in environments characterized by extremes — at very high or low temperatures, extreme pressure, high salinity, exceedingly acidic or basic environments, among other seemingly hostile conditions for life. By human standards, they are found in the most inhospitable places on Earth, including volcanoes, underneath polar ice, hydrothermal vents on the sea floor, and even in the presence of radiation and toxic waste.
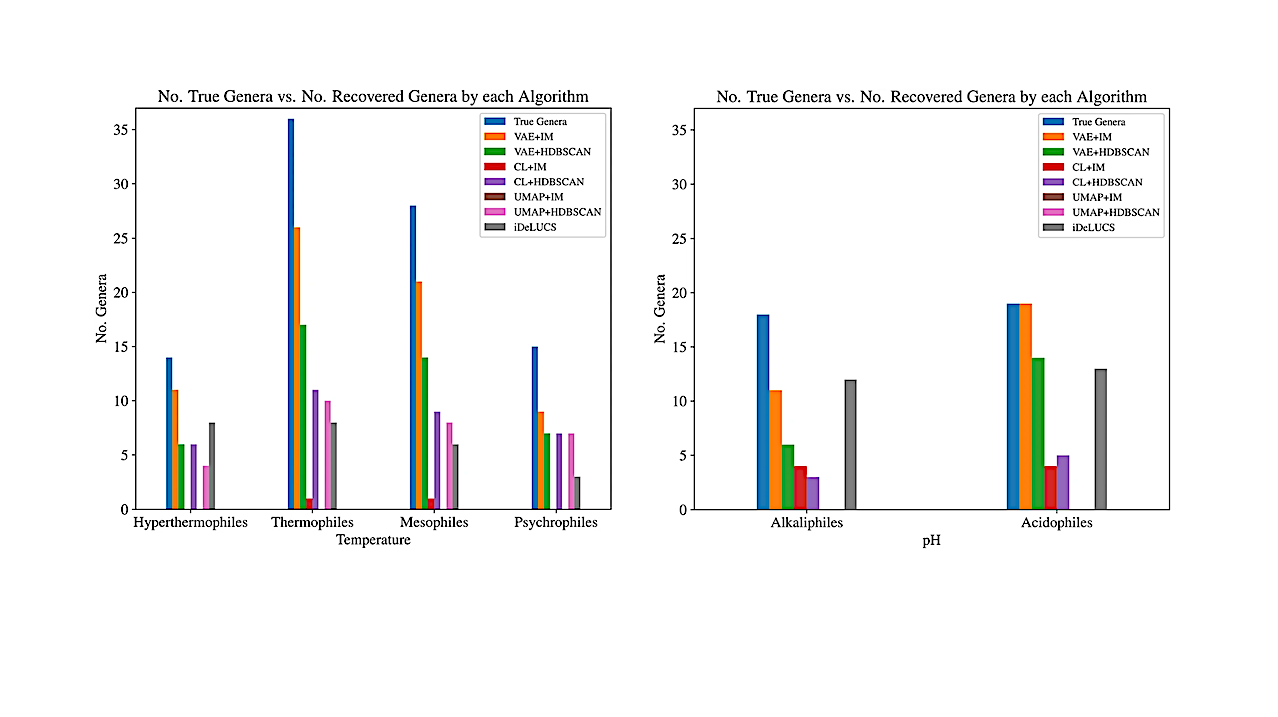
The research project evaluated a dataset of 700 microbial extremophile genomes using machine learning algorithms that looked for genomic similarities across them. Unexpectedly, the algorithms revealed that adaptations to extreme temperature or pH imprinted discernible patterns in the genomes of extremophiles, even though some belonged to different biological domains — microbes that are on entirely different limbs of the Tree of Life.
Such extremophiles are evolutionarily much more different from each other “than a polar bear is from a mushroom,” Kari said, adding that the detected genomic similarities therefore cannot be explained by ancestry. “That’s a fascinating insight, that an environmental signal exists, like a watermark, pervasive throughout the genome of some distantly related extremophiles that live in similar extreme environments, and that this environmental genomic signal can sometimes be stronger than even the genomic signal resulting from ancestry.”
“Our study has revealed, in some sense, a new dimension of the genome: The DNA of extremophiles contains, in addition to ancestry information, also information associated with the extreme environment where they live. This research could lay the groundwork for a “genomic observatory’’ to explore and discover other such signals within the genome. The results could potentially lead to a deeper understanding of the history of life on Earth.”
The research study, “Environment and taxonomy shape the genomic signature of prokaryotic extremophiles” (open access) co-authored with Pablo Millán Arias, Joseph Butler, Gurjit S. Randhawa, Maximillian P. M. Soltysiak, Kathleen A. Hill, and Kari, was recently published in the Nature journal Scientific Reports.
Astrobiology, genomics, evolution,