Self-Assembling Proteins Can Store Cellular “Memories”
As cells perform their everyday functions, they turn on a variety of genes and cellular pathways.
MIT engineers have now coaxed cells to inscribe the history of these events in a long protein chain that can be imaged using a light microscope.
Cells programmed to produce these chains continuously add building blocks that encode particular cellular events. Later, the ordered protein chains can be labeled with fluorescent molecules and read under a microscope, allowing researchers to reconstruct the timing of the events.
This technique could help shed light on the steps that underlie processes such as memory formation, response to drug treatment, and gene expression.
“There are a lot of changes that happen at organ or body scale, over hours to weeks, which cannot be tracked over time,” says Edward Boyden, the Y. Eva Tan Professor in Neurotechnology, a professor of biological engineering and brain and cognitive sciences at MIT, a Howard Hughes Medical Institute investigator, and a member of MIT’s McGovern Institute for Brain Research and Koch Institute for Integrative Cancer Research.
If the technique could be extended to work over longer time periods, it could also be used to study processes such as aging and disease progression, the researchers say.
Boyden is the senior author of the study, which appears today in Nature Biotechnology. Changyang Linghu, a former J. Douglas Tan Postdoctoral Fellow at the McGovern Institute, who is now an assistant professor at the University of Michigan, is the lead author of the paper.
Cellular history
Biological systems such as organs contain many different kinds of cells, all of which have distinctive functions. One way to study these functions is to image proteins, RNA, or other molecules inside the cells, which provide hints to what the cells are doing. However, most methods for doing this offer only a glimpse of a single moment in time, or don’t work well with very large populations of cells.
“Biological systems are often composed of a large number of different types of cells. For example, the human brain has 86 billion cells,” Linghu says. “To understand those kinds of biological systems, we need to observe physiological events over time in these large cell populations.”
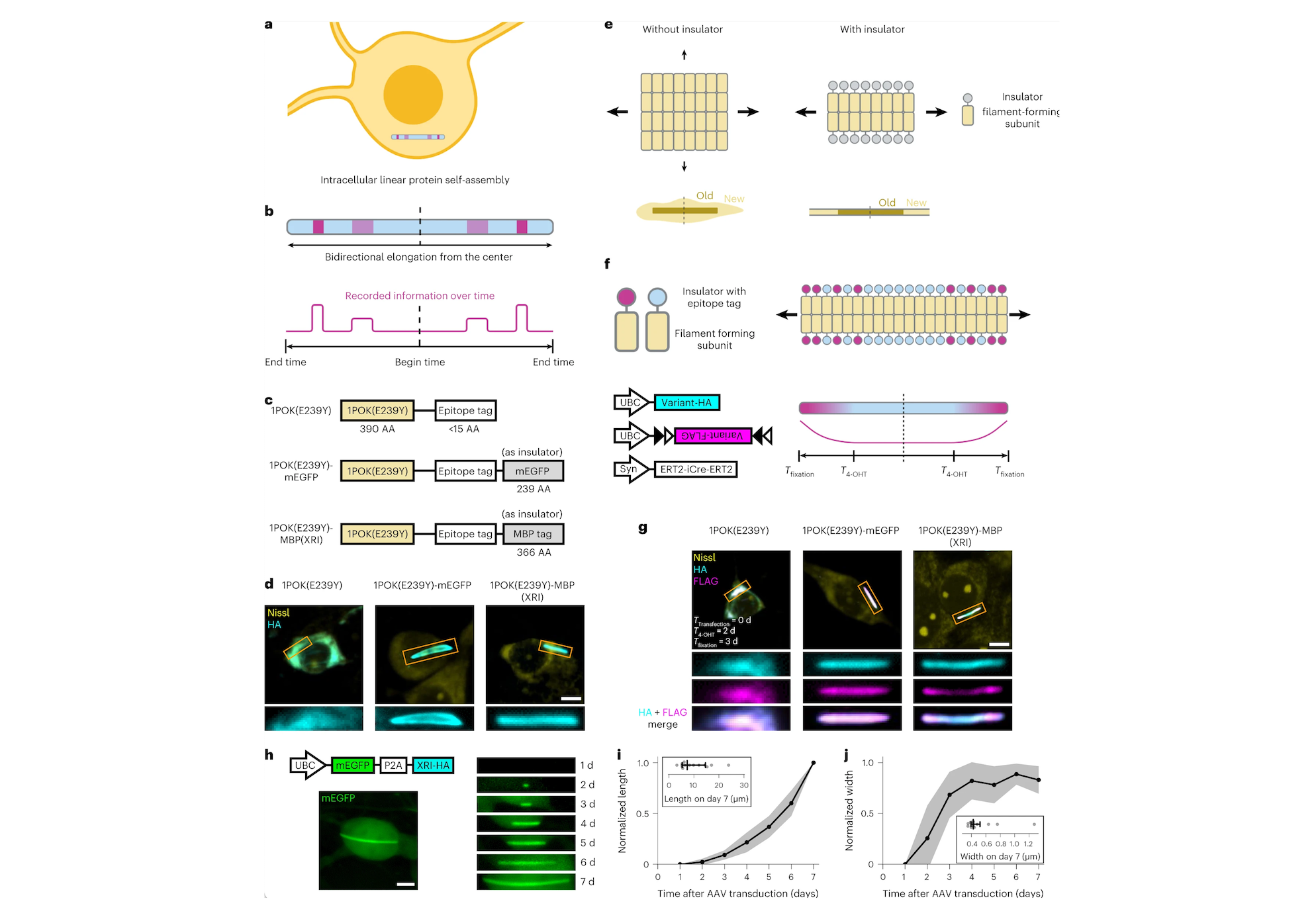
To achieve that, the research team came up with the idea of recording cellular events as a series of protein subunits that are continuously added to a chain. To create their chains, the researchers used engineered protein subunits, not normally found in living cells, that can self-assemble into long filaments.
The researchers designed a genetically encoded system in which one of these subunits is continuously produced inside cells, while the other is generated only when a specific event occurs. Each subunit also contains a very short peptide called an epitope tag — in this case, the researchers chose tags called HA and V5. Each of these tags can bind to a different fluorescent antibody, making it easy to visualize the tags later on and determine the sequence of the protein subunits.
For this study, the researchers made production of the V5-containing subunit contingent on the activation of a gene called c-fos, which is involved in encoding new memories. HA-tagged subunits make up most of the chain, but whenever the V5 tag shows up in the chain, that means that c-fos was activated during that time.
“We’re hoping to use this kind of protein self-assembly to record activity in every single cell,” Linghu says. “It’s not only a snapshot in time, but also records past history, just like how tree rings can permanently store information over time as the wood grows.”
Recording events
In this study, the researchers first used their system to record activation of c-fos in neurons growing in a lab dish. The c-fos gene was activated by chemically induced activation of the neurons, which caused the V5 subunit to be added to the protein chain.
To explore whether this approach could work in the brains of animals, the researchers programmed brain cells of mice to generate protein chains that would reveal when the animals were exposed to a particular drug. Later, the researchers were able to detect that exposure by preserving the tissue and analyzing it with a light microscope.
The researchers designed their system to be modular, so that different epitope tags can be swapped in, or different types of cellular events can be detected, including, in principle, cell division or activation of enzymes called protein kinases, which help control many cellular pathways.
The researchers also hope to extend the recording period that they can achieve. In this study, they recorded events for several days before imaging the tissue. There is a tradeoff between the amount of time that can be recorded and the time resolution, or frequency of event recording, because the length of the protein chain is limited by the size of the cell.
“The total amount of information it could store is fixed, but we could in principle slow down or increase the speed of the growth of the chain,” Linghu says. “If we want to record for a longer time, we could slow down the synthesis so that it will reach the size of the cell within, let’s say two weeks. In that way we could record longer, but with less time resolution.”
The researchers are also working on engineering the system so that it can record multiple types of events in the same chain, by increasing the number of different subunits that can be incorporated.
The research was funded by the Hock E. Tan and K. Lisa Yang Center for Autism Research, John Doerr, the National Institutes of Health, the National Science Foundation, the U.S. Army Research Office, and the Howard Hughes Medical Institute.
Recording of cellular physiological histories along optically readable self-assembling protein chains, Nature Biotechnology
Astrobiology