Autotrophic Biofilms Sustained By Deeply-sourced Groundwater Host Diverse CPR Bacteria Implicated In Sulfur And Hydrogen Metabolism
Candidate Phyla Radiation (CPR) bacteria are commonly detected yet enigmatic members of diverse microbial communities.
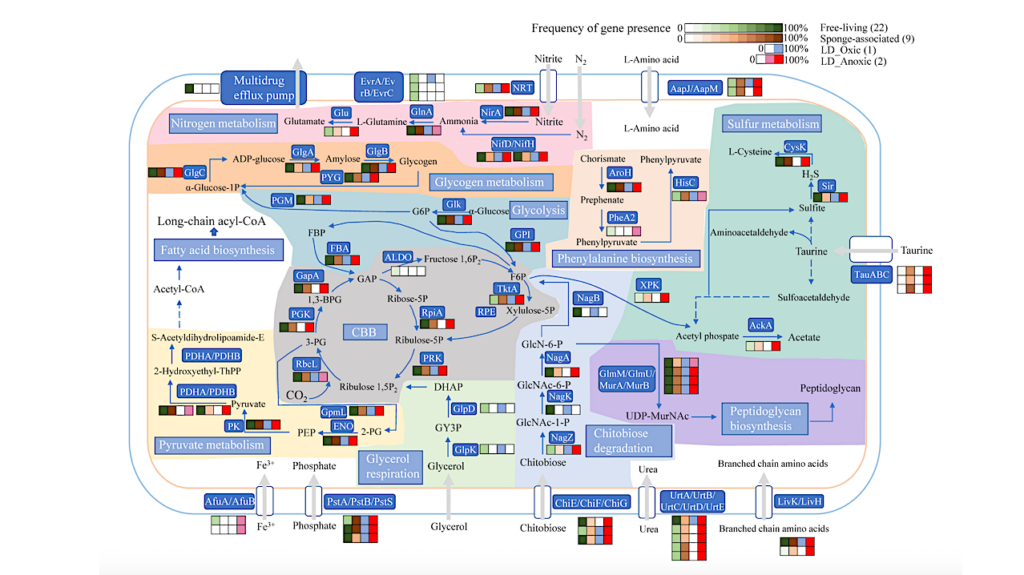
Their host associations, metabolic capabilities, and potential roles in biogeochemical cycles remain under-explored. We studied chemoautotrophically-based biofilms that host diverse CPR bacteria and grow in sulfide-rich springs using bulk geochemical analysis, genome-resolved metagenomics and scanning transmission x-ray microscopy (STXM) at room temperature and 87K. CPR-affiliated Gracilibacteria, Absconditabacteria, Saccharibacteria, Peregrinibacteria, Berkelbacteria, Microgenomates, and Parcubacteria are members of two biofilm communities dominated by chemolithotrophic sulfur-oxidizing bacteria including Thiothrix or Beggiatoa.
STXM imaging revealed ultra-small cells along the surfaces of filamentous bacteria that we interpret are CPR bacterial episymbionts. STXM and NEXAFS spectroscopy at carbon K and sulfur L2,3 edges show protein-encapsulated elemental sulfur spherical granules associated with filamentous bacteria, indicating that they are sulfur-oxidizers, likely Thiothrix. Berkelbacteria and Moranbacteria in the same biofilm sample are predicted to have a novel electron bifurcating group 3b [NiFe]-hydrogenase, putatively a sulfhydrogenase, potentially linked to sulfur metabolism via redox cofactors.
This complex could potentially underpin a symbiosis involving Berkelbacteria and/or Moranbacteria and filamentous sulfur-oxidizing bacteria such as Thiothrix that is based on cryptic sulfur cycling. One Doudnabacteria genome encodes adjacent sulfur dioxygenase and rhodanese genes that may convert thiosulfate to sulfite.
We find similar conserved genomic architecture associated with CPR bacteria from other sulfur-rich subsurface ecosystems. Our combined metagenomic, geochemical, spectromicroscopic and structural bioinformatics analyses link some CPR bacteria to sulfur-oxidizing Proteobacteria, likely Thiothrix, and indicate roles for CPR bacteria in sulfur and hydrogen cycling.
Competing Interest Statement
JFB is a co-founder of Metagenomi.
Luis E. Valentin Alvarado, Sirine C. Fakra, Alexander J. Probst, Jonathan R. Giska, Alexander L. Jaffe, Luke M. Oltrogge, View Jacob West-Roberts, Joel Rowland, Michael Manga, David F. Savage, Chris Greening, Brett J. Baker, Jillian F. Banfield
doi: https://doi.org/10.1101/2022.11.17.516901
https://www.eurekalert.org/news-releases/972799
Astrobiology